Frequently Asked Questions¶
| Author: | Kyle M. Douglass |
|---|---|
| Contact: | kyle.m.douglass@gmail.com |
| organization: | École Polytechnique Fédérale de Lausanne (EPFL) |
| revision: | $Revision: 5 $ |
| date: | 2017-01-24 |
| abstract: | This document answers frequently asked questions regarding B-Store, a lightweight data management system for single molecule localization microscopy (SMLM). |
Table of Contents
What is B-Store?¶
B-Store is a lightweight data management and analysis library for single molecule localization microscopy (SMLM). It serves two primary roles:
- To structure SMLM data inside a single, high performance filetype for fast and easy information retrieval and storage.
- To facilitate the analysis of high-throughput SMLM datasets.
What problem does B-Store solve?¶
High-throughput SMLM experiments can produce hundreds or even thousands of files containing multiple types of data (images, raw localizations, acquisition information, etc.). B-Store automatically sorts and stores this information in a datastore for rapid retrieval and analysis, removing any need to manually maintain the data yourself.
What are the design criteria for B-Store?¶
To realize these roles, B-Store is designed to meet these important criteria:
- Experimental datasets must be combined into a database-like structure that is easily readable by both humans and computers.
- Access and processing of data must be fast, regardless of the size of the dataset.
- Data provenance must be preserved throughout the organization and analysis pipeline.
- B-Store should not enforce standards that force scientists to adopt file formats, naming conventions, or software packages that differ from the ones they already use, except when it is absolutely necessary to achieve its roles.
- B-Store should be extensible to adapt to the changing needs of scientists using SMLM.
- Above all else, B-Store should make it easy to organize and document data and analysis pipelines to improve the reproducibility of SMLM experiments.
Of course, the changing needs of scientists means that B-Store will always be evolving to meet these criteria.
What doesn’t B-Store do?¶
B-Store is efficient and fast because its scope is limited to SMLM data organization and analysis. In particular, B-Store does not:
- Calculate localizations from raw images.
- Control microscopy hardware.
- Provide database-like storage for core facilities.
- Generate any data or results for you. (Sorry.)
Why don’t you use OME tools?¶
The Open Microscopy Environment (OME) is a wonderful set of software tools for working with bio-image data. In fact, the OME inspired this project in that B-Store emulates the OME model for archiving data, metadata, and analyses together in one abstract unit to improve reproducibility and communication of scientific results in SMLM.
In spite of this, we chose to develop tools independent of the OME for a few reasons. The OME was primarily designed for working with image data. SMLM data on the other hand is more heterogeneous than image data (localizations, drift correction, widefield images, etc.). Reworking parts of the OME to accomodate SMLM would therefore have been a significant undertaking on our part.
In addition, the OME database tool, OMERO, requires time for set up and maintenance. Many small labs doing SMLM may not be willing to invest the resources required for this. In contrast, B-Store is intended to be lightweight and require as little time for setup and maintenance as possible.
Some researchers in the SMLM community have expressed interest in extending the OME to SMLM, and we gladly welcome this effort. In the meanwhile, B-Store intends to satisfy the need for structured SMLM data.
How do I use B-Store?¶
B-Store is currently comprised of a set of functions, classes, and interfaces that are written in Python. You therefore can make B-Store datastores in any environment that runs Python code, including:
- The B-Store GUI
- Jupyter Notebooks
- IPython
- .py scripts
Once inside the datastore, the data may be accessed by any software that can read the HDF file format, including
and more.
Is there a GUI interface?¶
There is currently a lightweight GUI interface for building HDF datastores.
Can I still use B-Store if I don’t know Python?¶
If you don’t know Python, you can still use B-Store in a number of ways.
The easiest way is to use the GUI. After that, try exploring the Jupyter notebooks in the examples folder. Find an example that does what you want, then modify the relevant parts, such as file names. Then, simply run the notebook.
You may also wish to use B-Store’s datastore system, but not its analysis tools. In this case, you can use the notebooks to build your database, but access and analyze the data from the programming language of your choice, such as MATLAB. B-Store currently provides functionality for a datastore stored in an HDF file.
A third option is to call the Python code from within another language. Information for doing this in MATLAB may be found at the following link, though we have not yet tested this ourselves: http://www.mathworks.com/help/matlab/call-python-libraries.html
Of course, these approaches will only take you so far. Many parts of B-Store are meant to be customized to suit each scientist’s needs, and these customizations are most easily implemented in Python. Regardless, the largest amount of customization you will want to do will likely be to write a Parser. A Parser converts raw acquisition and localization data into a format that can pass through the datastore interface. If your programming language can call Python and the HDFDatastore object, then you can write the parser in the language of your choice and then pass the parsed data through these interfaces to build your database.
How do I contribute to or extend B-Store?¶
B-Store was designed to be extensible. If you have an idea, code, or even a comment about how to improve it, we would love to hear about it!
A great place to start contributing is by posting questions or comments to the B-Store mailing list.
Common extensions you would want to do are to write plugins that extend the Parser, Processor, or Reader classes, or write your own DatasetTypes. If you add your custom Python files to the ~/.bstore/bsplugins directory (%USERPROFILE%.bstore\bsplugins on Windows), B-Store will know to search this directory for imports.
A custom Parser that we use in our own lab may be found here: https://github.com/kmdouglass/bsplugins-leb
How do I add my custom code to the B-Store project?¶
If you want to modify the B-Store code, you can start by forking the repository on GitHub. According to GitHub’s documentation,
A fork is a copy of a repository. Forking a repository allows you to freely experiment with changes without affecting the original project.
After forking the repository, go ahead and make your changes, write some tests to be sure that your changes work like you expect them to, and then issue a pull request. The B-Store developers will review your suggested changes and, if they like them, will incorporate them into the B-Store project. With your permission your name will be added to the authors list.
For testing, B-Store uses the nose package. Type nosetests in the B-Store project root to run them. Test files are in the test files repository already mentioned. To run these successfully, set the __Path_To_Test_Data__ variable in bstore/config.py.
What language is B-Store written in?¶
B-Store is written in the Python programming language (version 3) and relies heavily on a datatype known as a DataFrame. DataFrames and their functionality are provided by the Pandas library and in many ways work like Excel spreadsheets but are much, much faster. Pandas is highly optimized and used extensively for both normal and big data analytics at companies and research institutions across the globe.
In addition to Pandas, B-Store implements features provided by numerous scientific, open source Python libraries like numpy and matplotlib. If you can’t do something in B-Store, you can likely still use these libraries to achieve what you want.
What is the logic of the B-Store datastore?¶
B-Store is designed to search specified directories on your computer for files associated with an SMLM experiment, such as those containing raw localizations and widefield images. These files are passed through a Parser, which converts them into a format suitable for insertion into a database. It does this by ensuring that the files satisfy the requirements of an interface known as a DatasetID. Data that implements this interface may pass into and out of the database; data that does not implement the interface cannot. You can think of the interface like a guard post at a government research facility. Only people with an ID badge for that facility (the interface) may enter. In principle, B-Store does not care about the data itself or the details of the database (HDF, SQL, etc.). At the moment, however, B-Store only supports databases contained in HDF files.
At the time this README file was written, the DatasetID of HDFDatastore consisted of the following properties:
- acquisition ID - integer identifying a specific acquisition
- prefix - a descriptive name for the acquisition, such as the cell type or condition
- datasetType - The type of data contained in the atom
- attribute of - For types that describe others, like localization metadata
- channel ID - the wavelength being imaged
- date ID - the date on which an acquisition was taken
- position ID - A single integer or integer pair identifying the position on the sample
- slice ID - An integer identifying the axial slice acquired
- replicate ID - An integer identifying the biological replicate that corresponds to this dataset.
The first four properties in bold are required; the last properties are optional.
There are three important advantages to enforcing an interface such as this.
- The computer will always know what kind of data it is working with and how to organize it.
- The format of the data that you generate in your experiments can be made independent of the datastore, so you can do whatever you want to it. The Parser ensures that it is in the right format only at the point of datastore insertion.
- The nature of the datastore and the types of data it can handle can grow and change in the future with minimal difficulty.
The logic of this interface is described graphically below. The raw data on top pass through the Parser and Readers objects and then into the datastore, where they are organized into acquisition groups. Each group is identified by a name called a prefix. Within the group, a dataset possesses an acquisition ID and a dataset type. An acqusition group is a set of datasets that were acquired during an experiment with the same prefix. A single dataset may optionally contain multiple fields of view (positions), wavelengths (channels), or axial slices. The database is therefore a collection of hierarchically arranged datasets, each belonging to a different acquisition group, and each uniquely identified by the conditions of the acquisition.
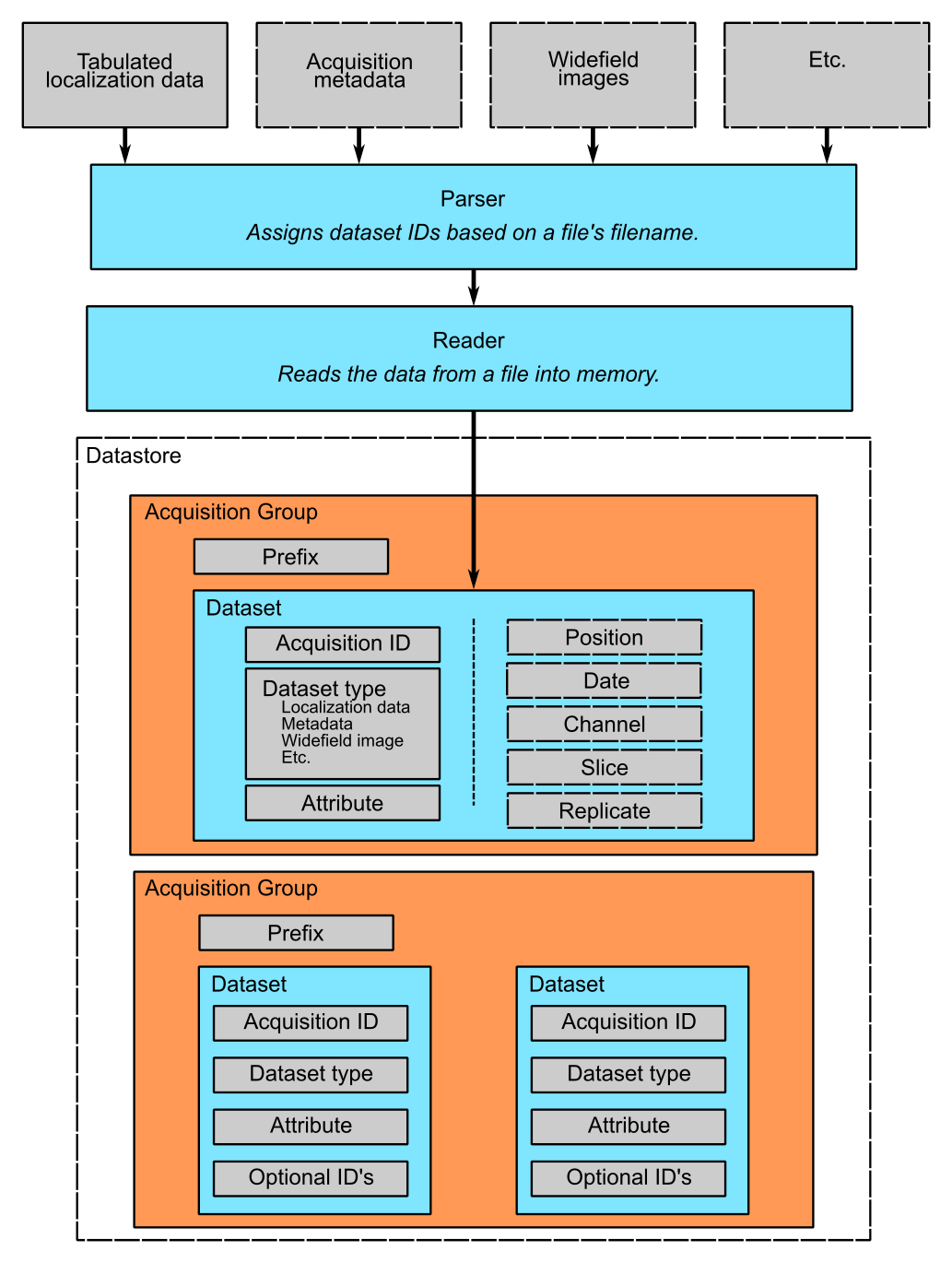
What is the logic behind the B-Store code?¶
The B-Store code base is divided into sixe separate modules:
- parsers
- database
- readers
- batch
- processors
- multiprocessors
In addition, functionality for each dataset type is specified in its own file in /bstore/datasetTypes/.
The first three modules, parsers, database, and readers, contain all the code for organizing SMLM datasets into a datastore. The last three modules, batch, processors, and multiprocessors, are primarily used for extracting data from B-Store databases and performing (semi-)automated analyses.
Parsers¶
A parser reads files from a SMLM acquisition and produces a Dataset–an object that can be inserted into a B-Store datastore. This object will have mandatory and possibly optional fields for uniquely identifying the data within the datastore.
Database¶
The database module contains code for building datastores from raw data. It relies on a parser for translating files into a format that it knows how to work with.
Readers¶
Readers understand how to read data from files generated by different sources, such as ThunderSTORM or RapidSTORM, and convert them into a common and internal Python data type. This internal representation is temporary and is used to next write this data to HDF.
Readers were introduced in version 1.1.0 and lay the groundwork for a more customizable interface in later versions.
Batch¶
The batch module contains routines for performing automated analyses with B-Store databases. It allows you to build simple analysis pipelines for extracting just the data you need from the datastore.
Processors¶
Processors are objects that take just a few parameters. When called, they accept a single argument (usually a Pandas DataFrame) as an input and produce an object of the same datatype as an output with its data having been modified.
Examples of processors include common SMLM analysis steps such as Filter, Merge, and Cluster.
Multiprocessors¶
Multiprocessors are similar to processors. They differ in that they take multiple inputs to produce an output. One multiprocessor is called OverlayClusters, which overlays clusters of localizations onto a widefield image for visual inspection and anotation of cluster analyses.
What testing framework is used by the B-Store developers?¶
Unit tests for B-Store are written as functions with utilities provided by Python’s nose package. Each module in B-Store has its own .py file containing these tests. They are stored in the bstore/tests and bstore/datasetTypes/tests folders in the B-Store root directory.
If you contribute to B-Store, we ask that you write unit tests for your code so that the developers can better understand what it’s supposed to do before merging it into the main project.
Gotcha’s¶
Spaces in column names¶
The library that B-Store uses to write to HDF files (PyTables) often has problems with spaces inside the names of DataFrame columns. We therefore recommend not using spaces. A workaround to this is to use the ConvertHeader processor to change column names during insertion into and retrieval from the database.
Widefield images¶
Grayscale¶
Widefield images are assumed to be grayscale. Unexpected behavior may result when attempting to place a color image into the database.
OME-XML¶
When reading metadata to determine the element_size_um attribute
of the HDF image_data, the OME-XML metadata tags PhysicalSizeX
and PhysicalSizeY will only be used if the corresponding units are
in microns. This means the PhysicalSizeXUnit and
PhysicalSizeYUnit must match the byte string \xc2\xb5m, which
is UTF-8 for the Greek letter “mu”, followed by the roman letter “m”.
If Micro-Manager (MM) metadata with pixel size information is present, then the OME-XML data will be ignored in favor of the MM metadata.
See the page on using B-Store in other software packages for more information.
What is single molecule localization microscopy (SMLM)?¶
SMLM is a suite of super-resolution fluorescence microscopy techniques for imaging microscopic structures (like cells and organelles) with resolutions below the diffraction limit of light. A number of SMLM techniques exist, such as fPALM, PALM, STORM, and PAINT. In these microscopies, fluorescent molecules are made to “blink” on and off. A final image or dataset is computed by recording the positions of every blink for a period of time and adding together all the positions in the end.
SMLM is a powerful tool for helping scientists understand biology and chemistry at nanometer length scales. It is particularly well-suited for structural biology and for tracking single fluorescent molecules in time.
A fantastic movie explaining how this works using the blinking lights of the Eiffel tower was created by Ricardo Henriques. You can watch it here: https://www.youtube.com/watch?v=RE70GuMCzww
What does the “B” stand for?¶
“Blink”